Overview of the tidyseurat package
Stefano Mangiola
2026-02-25
Source:vignettes/introduction.Rmd
introduction.RmdBrings Seurat to the tidyverse!
website: stemangiola.github.io/tidyseurat/
Please also have a look at
- tidyseurat for tidy single-cell RNA sequencing analysis
- tidySummarizedExperiment for tidy bulk RNA sequencing analysis
- tidybulk for tidy bulk RNA-seq analysis
- tidygate for adding custom gate information to your tibble
- tidyHeatmap for heatmaps produced with tidy principles

Introduction
tidyseurat provides a bridge between the Seurat single-cell package (Butler et al. 2018; Stuart et al. 2019) and the tidyverse (Wickham et al. 2019). It creates an invisible layer that enables viewing the Seurat object as a tidyverse tibble, and provides Seurat-compatible dplyr, tidyr, ggplot and plotly functions.
Functions/utilities available
| Seurat-compatible Functions | Description |
|---|---|
all |
| tidyverse Packages | Description |
|---|---|
dplyr |
All dplyr APIs like for any tibble |
tidyr |
All tidyr APIs like for any tibble |
ggplot2 |
ggplot like for any tibble |
plotly |
plot_ly like for any tibble |
| Utilities | Description |
|---|---|
tidy |
Add tidyseurat invisible layer over a Seurat
object |
as_tibble |
Convert cell-wise information to a tbl_df
|
join_features |
Add feature-wise information, returns a tbl_df
|
aggregate_cells |
Aggregate cell gene-transcription abundance as pseudobulk tissue |
Installation
From CRAN
install.packages("tidyseurat")From Github (development)
devtools::install_github("stemangiola/tidyseurat")Create tidyseurat, the best of both worlds!
This is a seurat object but it is evaluated as tibble. So it is fully compatible both with Seurat and tidyverse APIs.
pbmc_small = SeuratObject::pbmc_smallIt looks like a tibble
pbmc_small## # A Seurat-tibble abstraction: 80 × 15
## # Features=230 | Cells=80 | Active assay=RNA | Assays=RNA
## .cell orig.ident nCount_RNA nFeature_RNA RNA_snn_res.0.8 letter.idents groups
## <chr> <fct> <dbl> <int> <fct> <fct> <chr>
## 1 ATGC… SeuratPro… 70 47 0 A g2
## 2 CATG… SeuratPro… 85 52 0 A g1
## 3 GAAC… SeuratPro… 87 50 1 B g2
## 4 TGAC… SeuratPro… 127 56 0 A g2
## 5 AGTC… SeuratPro… 173 53 0 A g2
## 6 TCTG… SeuratPro… 70 48 0 A g1
## 7 TGGT… SeuratPro… 64 36 0 A g1
## 8 GCAG… SeuratPro… 72 45 0 A g1
## 9 GATA… SeuratPro… 52 36 0 A g1
## 10 AATG… SeuratPro… 100 41 0 A g1
## # ℹ 70 more rows
## # ℹ 8 more variables: RNA_snn_res.1 <fct>, PC_1 <dbl>, PC_2 <dbl>, PC_3 <dbl>,
## # PC_4 <dbl>, PC_5 <dbl>, tSNE_1 <dbl>, tSNE_2 <dbl>But it is a Seurat object after all
pbmc_small@assays## $RNA
## Assay data with 230 features for 80 cells
## Top 10 variable features:
## PPBP, IGLL5, VDAC3, CD1C, AKR1C3, PF4, MYL9, GNLY, TREML1, CA2Preliminary plots
Set colours and theme for plots.
# Use colourblind-friendly colours
friendly_cols <- c("#88CCEE", "#CC6677", "#DDCC77", "#117733", "#332288", "#AA4499", "#44AA99", "#999933", "#882255", "#661100", "#6699CC")
# Set theme
my_theme <-
list(
scale_fill_manual(values = friendly_cols),
scale_color_manual(values = friendly_cols),
theme_bw() +
theme(
panel.border = element_blank(),
axis.line = element_line(),
panel.grid.major = element_line(size = 0.2),
panel.grid.minor = element_line(size = 0.1),
text = element_text(size = 12),
legend.position = "bottom",
aspect.ratio = 1,
strip.background = element_blank(),
axis.title.x = element_text(margin = margin(t = 10, r = 10, b = 10, l = 10)),
axis.title.y = element_text(margin = margin(t = 10, r = 10, b = 10, l = 10))
)
)We can treat pbmc_small effectively as a normal tibble
for plotting.
Here we plot number of features per cell.
pbmc_small %>%
ggplot(aes(nFeature_RNA, fill = groups)) +
geom_histogram() +
my_theme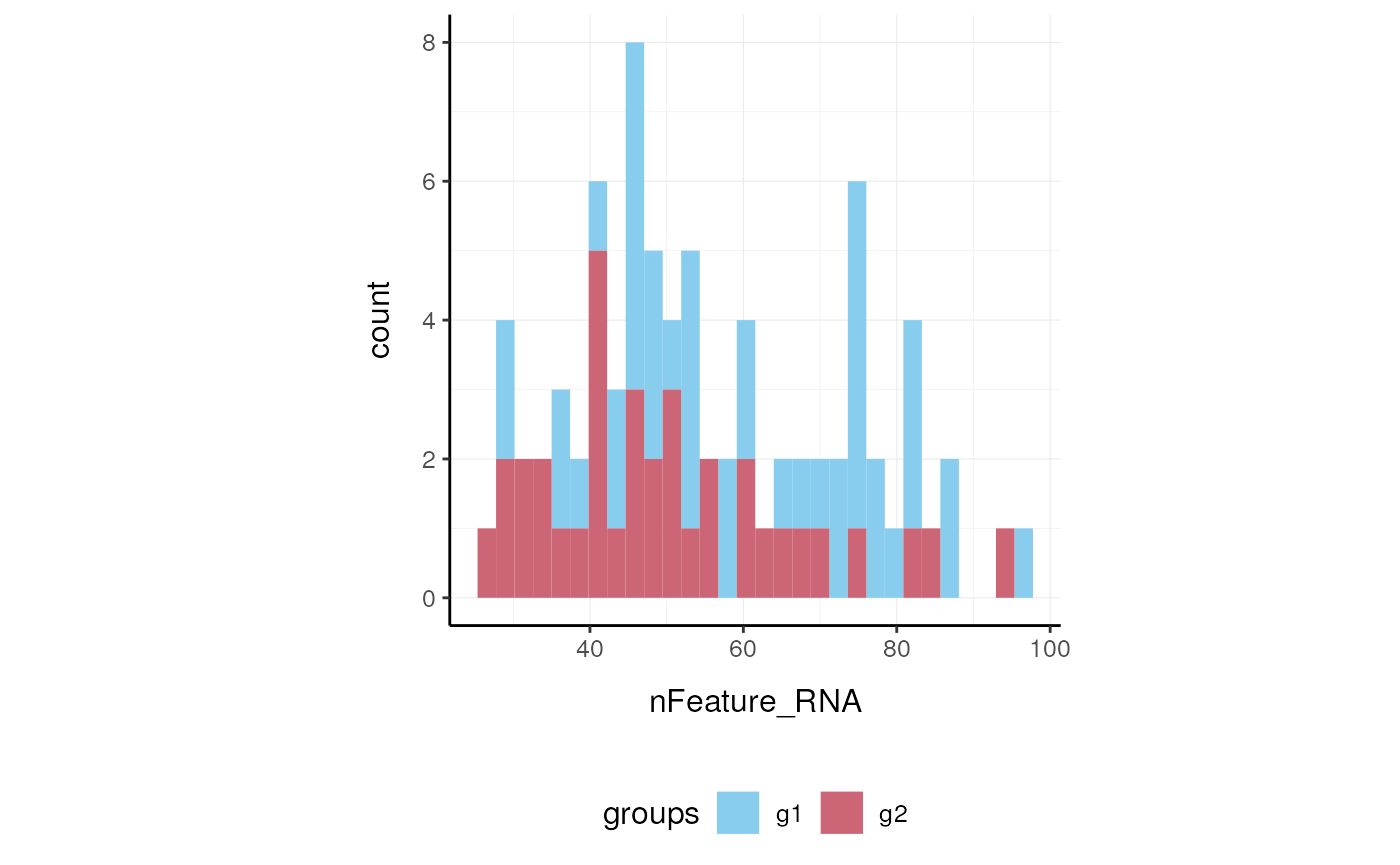
Here we plot total features per cell.
pbmc_small %>%
ggplot(aes(groups, nCount_RNA, fill = groups)) +
geom_boxplot(outlier.shape = NA) +
geom_jitter(width = 0.1) +
my_theme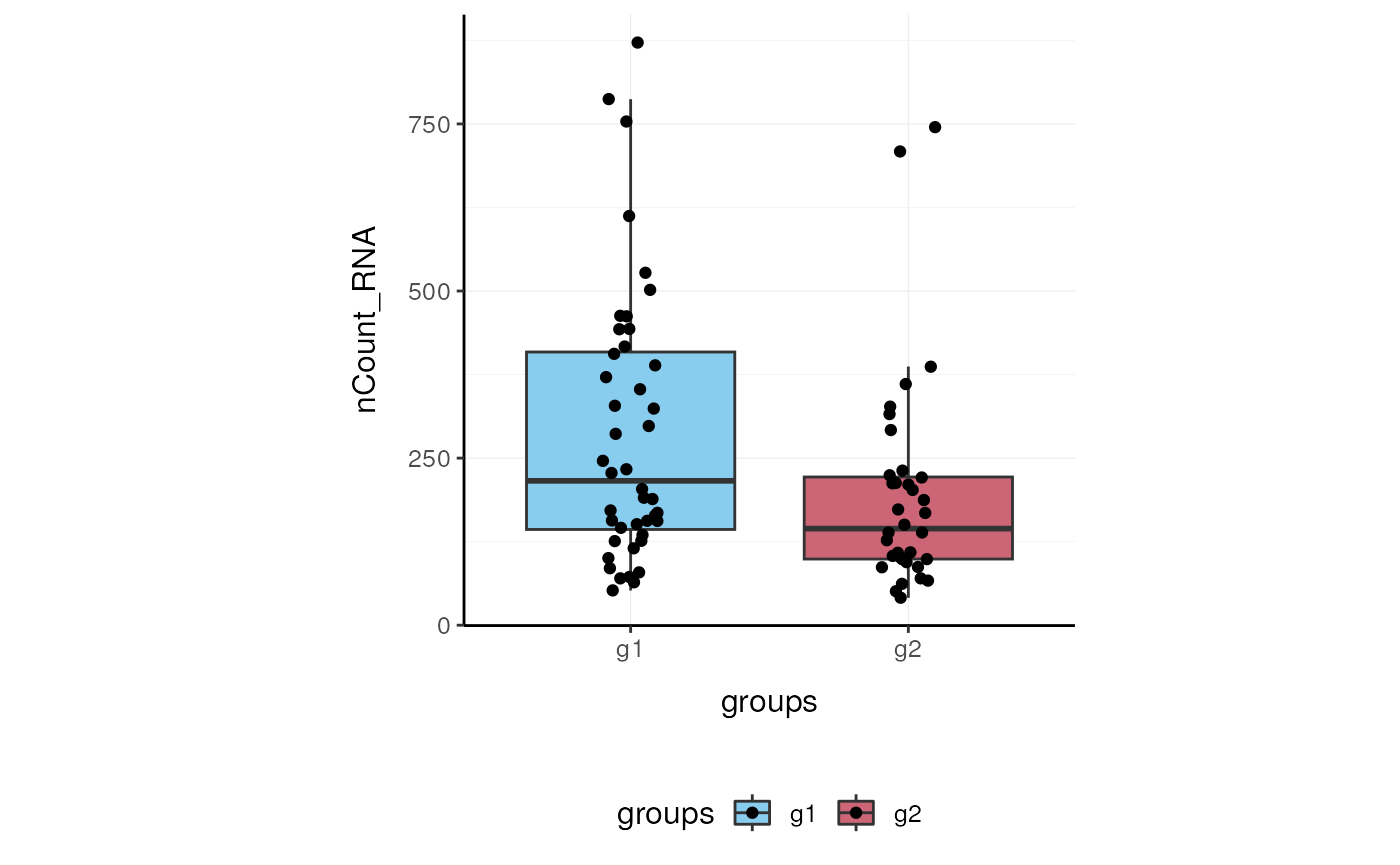
Here we plot abundance of two features for each group.
pbmc_small %>%
join_features(features = c("HLA-DRA", "LYZ"), shape = "long") %>%
ggplot(aes(groups, .abundance_RNA + 1, fill = groups)) +
geom_boxplot(outlier.shape = NA) +
geom_jitter(aes(size = nCount_RNA), alpha = 0.5, width = 0.2) +
scale_y_log10() +
my_theme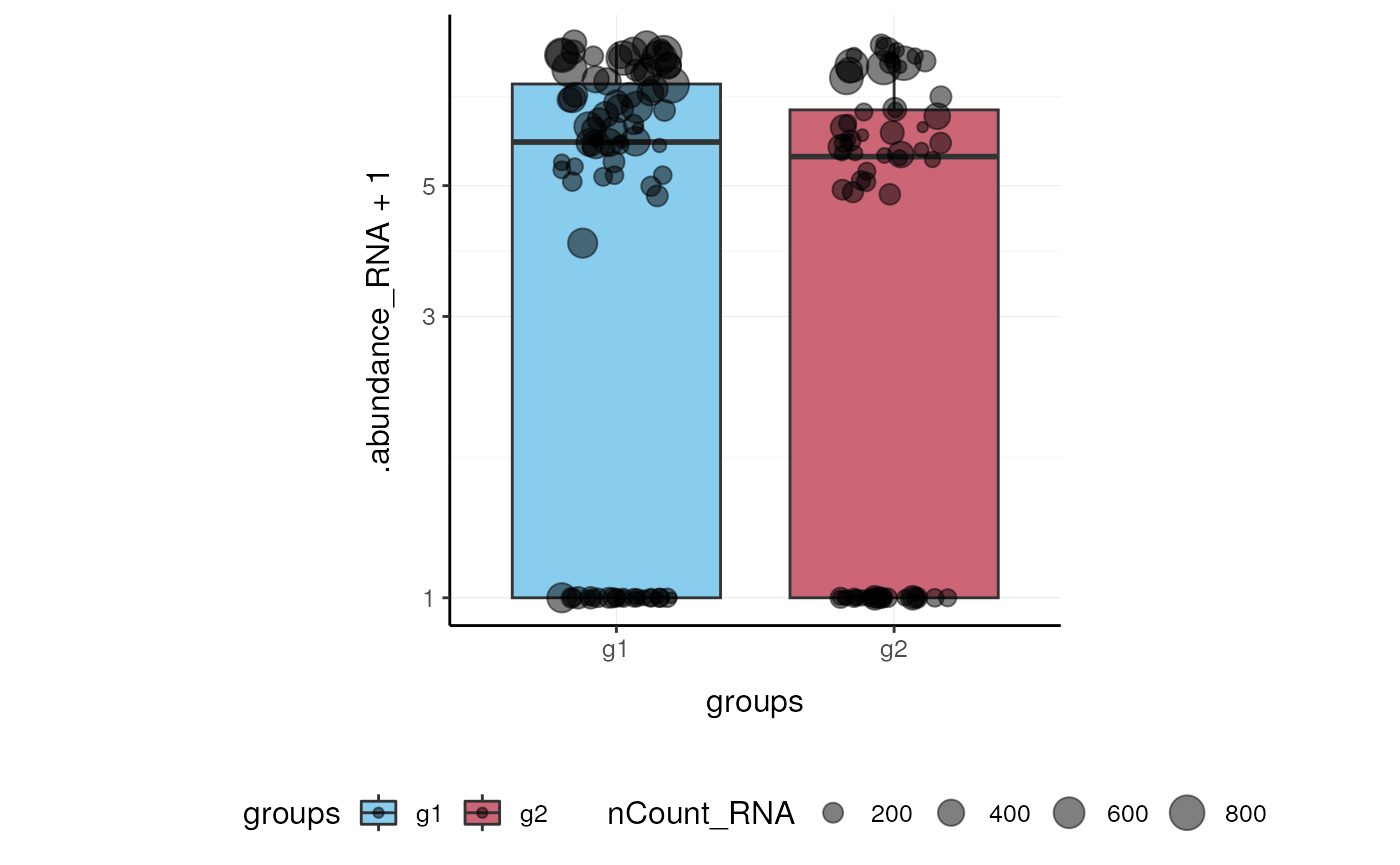
Preprocess the dataset
Also you can treat the object as Seurat object and proceed with data processing.
pbmc_small_pca <-
pbmc_small %>%
SCTransform(verbose = FALSE) %>%
FindVariableFeatures(verbose = FALSE) %>%
RunPCA(verbose = FALSE)
pbmc_small_pca## # A Seurat-tibble abstraction: 80 × 17
## # Features=220 | Cells=80 | Active assay=SCT | Assays=RNA, SCT
## .cell orig.ident nCount_RNA nFeature_RNA RNA_snn_res.0.8 letter.idents groups
## <chr> <fct> <dbl> <int> <fct> <fct> <chr>
## 1 ATGC… SeuratPro… 70 47 0 A g2
## 2 CATG… SeuratPro… 85 52 0 A g1
## 3 GAAC… SeuratPro… 87 50 1 B g2
## 4 TGAC… SeuratPro… 127 56 0 A g2
## 5 AGTC… SeuratPro… 173 53 0 A g2
## 6 TCTG… SeuratPro… 70 48 0 A g1
## 7 TGGT… SeuratPro… 64 36 0 A g1
## 8 GCAG… SeuratPro… 72 45 0 A g1
## 9 GATA… SeuratPro… 52 36 0 A g1
## 10 AATG… SeuratPro… 100 41 0 A g1
## # ℹ 70 more rows
## # ℹ 10 more variables: RNA_snn_res.1 <fct>, nCount_SCT <dbl>,
## # nFeature_SCT <int>, PC_1 <dbl>, PC_2 <dbl>, PC_3 <dbl>, PC_4 <dbl>,
## # PC_5 <dbl>, tSNE_1 <dbl>, tSNE_2 <dbl>If a tool is not included in the tidyseurat collection, we can use
as_tibble to permanently convert tidyseurat
into tibble.
pbmc_small_pca %>%
as_tibble() %>%
select(contains("PC"), everything()) %>%
GGally::ggpairs(columns = 1:5, ggplot2::aes(colour = groups)) +
my_theme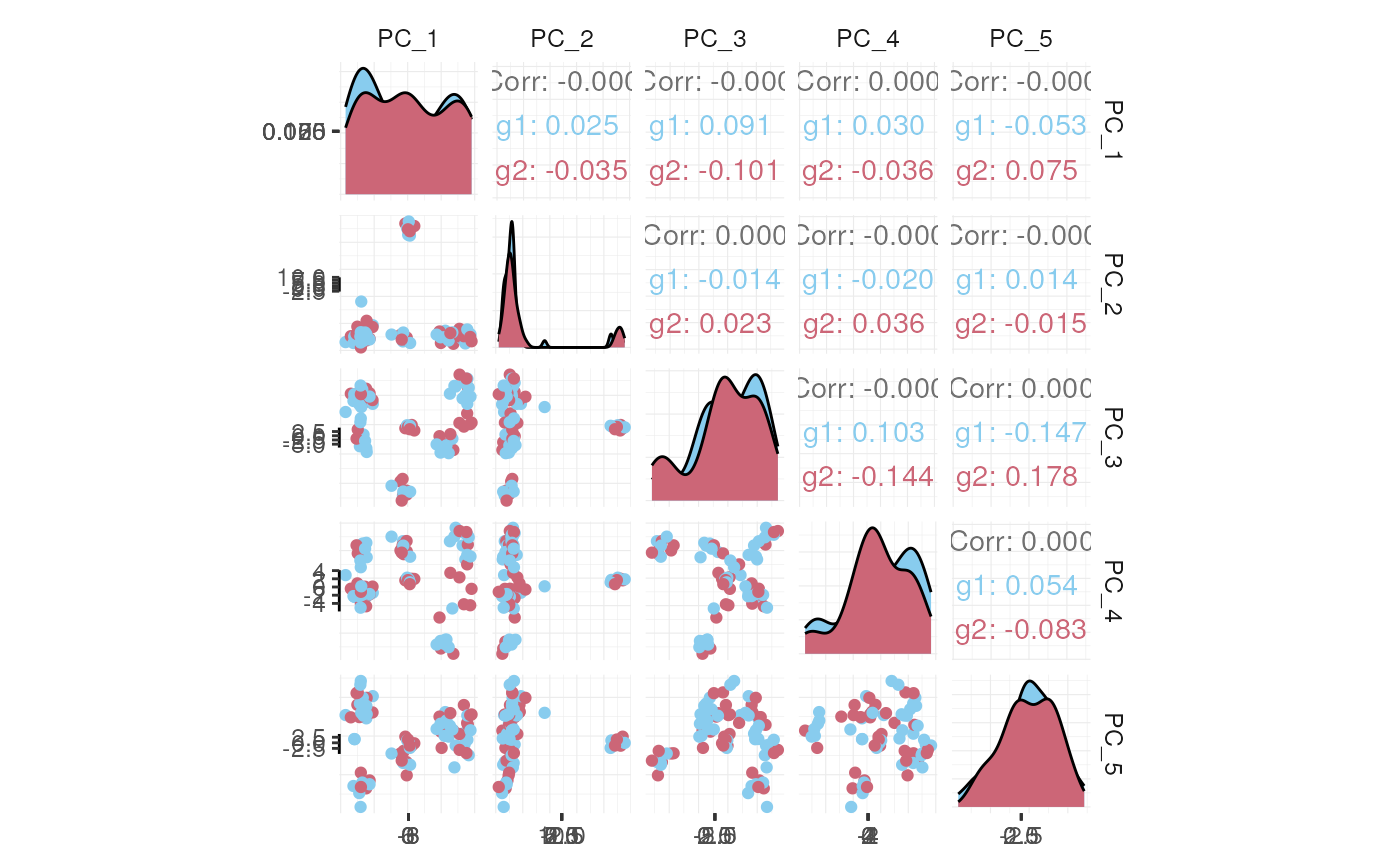
Identify clusters
We proceed with cluster identification with Seurat.
pbmc_small_cluster <-
pbmc_small_pca %>%
FindNeighbors(verbose = FALSE) %>%
FindClusters(method = "igraph", verbose = FALSE)
pbmc_small_cluster## # A Seurat-tibble abstraction: 80 × 19
## # Features=220 | Cells=80 | Active assay=SCT | Assays=RNA, SCT
## .cell orig.ident nCount_RNA nFeature_RNA RNA_snn_res.0.8 letter.idents groups
## <chr> <fct> <dbl> <int> <fct> <fct> <chr>
## 1 ATGC… SeuratPro… 70 47 0 A g2
## 2 CATG… SeuratPro… 85 52 0 A g1
## 3 GAAC… SeuratPro… 87 50 1 B g2
## 4 TGAC… SeuratPro… 127 56 0 A g2
## 5 AGTC… SeuratPro… 173 53 0 A g2
## 6 TCTG… SeuratPro… 70 48 0 A g1
## 7 TGGT… SeuratPro… 64 36 0 A g1
## 8 GCAG… SeuratPro… 72 45 0 A g1
## 9 GATA… SeuratPro… 52 36 0 A g1
## 10 AATG… SeuratPro… 100 41 0 A g1
## # ℹ 70 more rows
## # ℹ 12 more variables: RNA_snn_res.1 <fct>, nCount_SCT <dbl>,
## # nFeature_SCT <int>, SCT_snn_res.0.8 <fct>, seurat_clusters <fct>,
## # PC_1 <dbl>, PC_2 <dbl>, PC_3 <dbl>, PC_4 <dbl>, PC_5 <dbl>, tSNE_1 <dbl>,
## # tSNE_2 <dbl>Now we can interrogate the object as if it was a regular tibble data frame.
## # A tibble: 6 × 3
## groups seurat_clusters n
## <chr> <fct> <int>
## 1 g1 0 23
## 2 g1 1 17
## 3 g1 2 4
## 4 g2 0 17
## 5 g2 1 13
## 6 g2 2 6We can identify cluster markers using Seurat.
Reduce dimensions
We can calculate the first 3 UMAP dimensions using the Seurat framework.
pbmc_small_UMAP <-
pbmc_small_cluster %>%
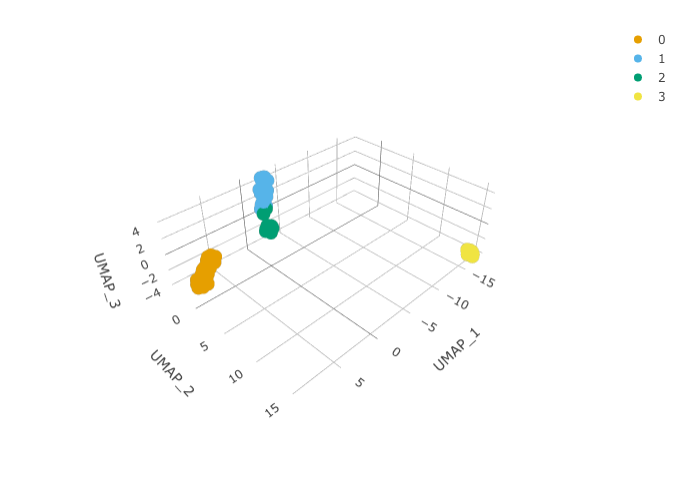
RunUMAP(reduction = "pca", dims = 1:15, n.components = 3L)And we can plot them using 3D plot using plotly.
pbmc_small_UMAP %>%
plot_ly(
x = ~`UMAP_1`,
y = ~`UMAP_2`,
z = ~`UMAP_3`,
color = ~seurat_clusters,
colors = friendly_cols[1:4]
)
Cell type prediction
We can infer cell type identities using SingleR (Aran et al. 2019) and manipulate the output using tidyverse.
# Get cell type reference data
blueprint <- celldex::BlueprintEncodeData()
# Infer cell identities
cell_type_df <-
GetAssayData(pbmc_small_UMAP, slot = 'counts', assay = "SCT") %>%
log1p() %>%
Matrix::Matrix(sparse = TRUE) %>%
SingleR::SingleR(
ref = blueprint,
labels = blueprint$label.main,
method = "single"
) %>%
as.data.frame() %>%
as_tibble(rownames = "cell") %>%
select(cell, first.labels)
# Join UMAP and cell type info
pbmc_small_cell_type <-
pbmc_small_UMAP %>%
left_join(cell_type_df, by = "cell")
# Reorder columns
pbmc_small_cell_type %>%
select(cell, first.labels, everything())We can easily summarise the results. For example, we can see how cell type classification overlaps with cluster classification.
We can easily reshape the data for building information-rich faceted plots.
pbmc_small_cell_type %>%
# Reshape and add classifier column
pivot_longer(
cols = c(seurat_clusters, first.labels),
names_to = "classifier", values_to = "label"
) %>%
# UMAP plots for cell type and cluster
ggplot(aes(UMAP_1, UMAP_2, color = label)) +
geom_point() +
facet_wrap(~classifier) +
my_themeWe can easily plot gene correlation per cell category, adding multi-layer annotations.
pbmc_small_cell_type %>%
# Add some mitochondrial abundance values
mutate(mitochondrial = rnorm(n())) %>%
# Plot correlation
join_features(features = c("CST3", "LYZ"), shape = "wide") %>%
ggplot(aes(CST3 + 1, LYZ + 1, color = groups, size = mitochondrial)) +
geom_point() +
facet_wrap(~first.labels, scales = "free") +
scale_x_log10() +
scale_y_log10() +
my_themeNested analyses
A powerful tool we can use with tidyseurat is nest. We
can easily perform independent analyses on subsets of the dataset. First
we classify cell types in lymphoid and myeloid; then, nest based on the
new classification
pbmc_small_nested <-
pbmc_small_cell_type %>%
filter(first.labels != "Erythrocytes") %>%
mutate(cell_class = if_else(`first.labels` %in% c("Macrophages", "Monocytes"), "myeloid", "lymphoid")) %>%
nest(data = -cell_class)
pbmc_small_nestedNow we can independently for the lymphoid and myeloid subsets (i) find variable features, (ii) reduce dimensions, and (iii) cluster using both tidyverse and Seurat seamlessly.
pbmc_small_nested_reanalysed <-
pbmc_small_nested %>%
mutate(data = map(
data, ~ .x %>%
FindVariableFeatures(verbose = FALSE) %>%
RunPCA(npcs = 10, verbose = FALSE) %>%
FindNeighbors(verbose = FALSE) %>%
FindClusters(method = "igraph", verbose = FALSE) %>%
RunUMAP(reduction = "pca", dims = 1:10, n.components = 3L, verbose = FALSE)
))
pbmc_small_nested_reanalysedNow we can unnest and plot the new classification.
pbmc_small_nested_reanalysed %>%
# Convert to tibble otherwise Seurat drops reduced dimensions when unifying data sets.
mutate(data = map(data, ~ .x %>% as_tibble())) %>%
unnest(data) %>%
# Define unique clusters
unite("cluster", c(cell_class, seurat_clusters), remove = FALSE) %>%
# Plotting
ggplot(aes(UMAP_1, UMAP_2, color = cluster)) +
geom_point() +
facet_wrap(~cell_class) +
my_themeAggregating cells
Sometimes, it is necessary to aggregate the gene-transcript abundance from a group of cells into a single value. For example, when comparing groups of cells across different samples with fixed-effect models.
In tidyseurat, cell aggregation can be achieved using the
aggregate_cells function.
pbmc_small %>%
aggregate_cells(groups, assays = "RNA")Session Info
## R Under development (unstable) (2026-02-22 r89452)
## Platform: x86_64-pc-linux-gnu
## Running under: Ubuntu 24.04.4 LTS
##
## Matrix products: default
## BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
## LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
##
## locale:
## [1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
## [3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
## [5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
## [7] LC_PAPER=en_US.UTF-8 LC_NAME=C
## [9] LC_ADDRESS=C LC_TELEPHONE=C
## [11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
##
## time zone: UTC
## tzcode source: system (glibc)
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] future_1.69.0 tidyseurat_0.8.9 ttservice_0.5.3 Seurat_5.4.0
## [5] SeuratObject_5.3.0 sp_2.2-1 ggplot2_4.0.2 magrittr_2.0.4
## [9] purrr_1.2.1 tidyr_1.3.2 dplyr_1.2.0 knitr_1.51
##
## loaded via a namespace (and not attached):
## [1] deldir_2.0-4 pbapply_1.7-4 gridExtra_2.3
## [4] rlang_1.1.7 RcppAnnoy_0.0.23 otel_0.2.0
## [7] spatstat.geom_3.7-0 matrixStats_1.5.0 ggridges_0.5.7
## [10] compiler_4.6.0 png_0.1-8 systemfonts_1.3.1
## [13] vctrs_0.7.1 reshape2_1.4.5 stringr_1.6.0
## [16] pkgconfig_2.0.3 fastmap_1.2.0 labeling_0.4.3
## [19] utf8_1.2.6 promises_1.5.0 rmarkdown_2.30
## [22] ragg_1.5.0 xfun_0.56 cachem_1.1.0
## [25] jsonlite_2.0.0 goftest_1.2-3 later_1.4.7
## [28] spatstat.utils_3.2-1 irlba_2.3.7 parallel_4.6.0
## [31] cluster_2.1.8.2 R6_2.6.1 ica_1.0-3
## [34] spatstat.data_3.1-9 bslib_0.10.0 stringi_1.8.7
## [37] RColorBrewer_1.1-3 GGally_2.4.0 reticulate_1.45.0
## [40] spatstat.univar_3.1-6 parallelly_1.46.1 lmtest_0.9-40
## [43] jquerylib_0.1.4 scattermore_1.2 Rcpp_1.1.1
## [46] tensor_1.5.1 future.apply_1.20.2 zoo_1.8-15
## [49] sctransform_0.4.3 httpuv_1.6.16 Matrix_1.7-4
## [52] splines_4.6.0 igraph_2.2.2 tidyselect_1.2.1
## [55] abind_1.4-8 yaml_2.3.12 spatstat.random_3.4-4
## [58] spatstat.explore_3.7-0 codetools_0.2-20 miniUI_0.1.2
## [61] listenv_0.10.0 lattice_0.22-9 tibble_3.3.1
## [64] plyr_1.8.9 shiny_1.13.0 withr_3.0.2
## [67] S7_0.2.1 ROCR_1.0-12 evaluate_1.0.5
## [70] Rtsne_0.17 fastDummies_1.7.5 desc_1.4.3
## [73] survival_3.8-6 ggstats_0.12.0 polyclip_1.10-7
## [76] fitdistrplus_1.2-6 pillar_1.11.1 KernSmooth_2.23-26
## [79] plotly_4.12.0 generics_0.1.4 RcppHNSW_0.6.0
## [82] scales_1.4.0 globals_0.19.0 xtable_1.8-8
## [85] glue_1.8.0 lazyeval_0.2.2 tools_4.6.0
## [88] data.table_1.18.2.1 RSpectra_0.16-2 RANN_2.6.2
## [91] fs_1.6.6 dotCall64_1.2 cowplot_1.2.0
## [94] grid_4.6.0 nlme_3.1-168 patchwork_1.3.2
## [97] cli_3.6.5 spatstat.sparse_3.1-0 textshaping_1.0.4
## [100] fansi_1.0.7 spam_2.11-3 viridisLite_0.4.3
## [103] uwot_0.2.4 gtable_0.3.6 sass_0.4.10
## [106] digest_0.6.39 progressr_0.18.0 ggrepel_0.9.7
## [109] htmlwidgets_1.6.4 farver_2.1.2 htmltools_0.5.9
## [112] pkgdown_2.2.0 lifecycle_1.0.5 httr_1.4.8
## [115] mime_0.13 MASS_7.3-65